Final year students Annette Strege (graduated 2013) and Sorrell Bunting (graduated 2012), who both did lab projects with Dr Bernadette Byrne, were recently delighted to see their research work published in the scientific journal PLOS ONE. Annette tells the story:

During my final year project in 2013 I investigated the functional effects of introducing thermostabilising point mutations into a G-Protein Coupled Receptor (GPCR) in Dr. Bernadette Byrne’s lab. GPCRs are important proteins to study, because they are the most popular type of drug target, representing around 50% of all drug targets. However, these membrane receptors are also very challenging to study because they are very flexible and hydrophobic. In recent years, significant progress has been made in crystallising and obtaining high-resolution structural data for GPCRs. This is predominantly due to approaches that have been developed to stabilize the GPCR molecules prior to crystallisation. One very successful method for doing this is by introducing a small number of thermostabilising point mutations.
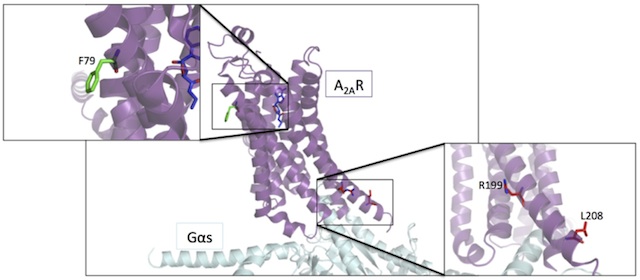
Although such approaches have been extremely successful for obtaining GPCR structures, the effects of the mutations on receptor function are not fully understood. While ligand binding is generally assessed quite widely, the activity of the pathways downstream of the receptor is rarely quantified. For my final year project, my task was therefore to use a functional assay to characterize the activity downstream of 30 mutants of the Adenosine A2A receptor (A2AR). These 30 mutants were based on a single thermostabilised variant of the A2AR, called Rag23. Together the 30 mutants we tested contained all combinations of the five thermostabilising point mutations found in Rag23.
The results of this project clearly showed that each of the five point mutations had a detectable effect on the activity profile of the A2AR. In addition, these effects were additive when individual mutations were combined. Furthermore, the existing crystal structures of the A2AR and other GPCRs provide a good foundation for explaining why we observe certain changes in function upon mutating certain structural regions.
I really enjoyed this project, because although 10 weeks is not a lot of time to get a good amount of work done, everything was so well organised when I started my work that producing good results was very quick. The mutants had been generated, the assay had been tested, and my task was simply to follow the protocol, run just over 60 assays, analyse the data, find the pattern, and explain why this pattern is observed based on structural data available in the literature.
I knew that the project had gone well and was expecting a first, but I never expected to get the highest mark out of everyone in my course. I was even more surprised and excited when I was told that the work would get published. Although writing, submission, corrections, re-submission, and so on was a rather long process that took more than 6 months, eventually going to the Library to print out my own paper felt amazing!