A new study carried out by Peter Sarkies (Transgenerational Epigenetic Inheritance & Evolution) in collaboration with Eric Miska (Gurdon Institute, Cambridge) reveals astonishing insights into the evolution of transposon-silencing mechanisms in nematode worms. Transposons are “selfish” DNA pieces that insert themselves into the genome. Like a computer virus, they copy themselves and proliferate, which can disrupt essential gene functions. Because transposons are so disruptive, there is huge selective pressure on organisms to silence them and stop them spreading. Organisms have evolved ingenious ways to suppress transposon activity, especially in the reproductive cells, where a transposition event affects subsequent generations. The front line of defence against transposons in most animals, from nematode worms to humans, are tiny sequences of RNA, known as Piwi interacting small RNAs (piRNAs). These piRNAs patrol the genome, seeking out and controlling transposons.
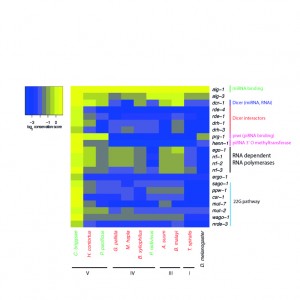 The phylum-wide study, published this week in PLOS Biology, sheds light on a highly dynamic evolutionary history, which was completely unexpected, whereby this broadly conserved transposon-silencing system has been lost in nematodes on several occasions. The study shows that piRNAs have been completely lost in four of the five nematode groups, or clades. Only Clade V, to which the lab model Caenorhabditis elegans belongs, use the piRNA pathway. In the absence of piRNAs, nematodes use a diversity of other mechanisms to control transposons. The authors suggest that the piRNA pathway was present in the most ancestral ur-nematode, but then independently lost in other nematode lineages.
The phylum-wide study, published this week in PLOS Biology, sheds light on a highly dynamic evolutionary history, which was completely unexpected, whereby this broadly conserved transposon-silencing system has been lost in nematodes on several occasions. The study shows that piRNAs have been completely lost in four of the five nematode groups, or clades. Only Clade V, to which the lab model Caenorhabditis elegans belongs, use the piRNA pathway. In the absence of piRNAs, nematodes use a diversity of other mechanisms to control transposons. The authors suggest that the piRNA pathway was present in the most ancestral ur-nematode, but then independently lost in other nematode lineages.
However, nematodes without this piRNA pathway are not riddled with transposons, but have evolved two other pathways that control transposons in nematodes. One is a novel transposon-silencing pathway known as 22G-RNAs. This was found in three clades (Clades III, IV and V). The other is an ancient pathway, dependent on RNA-directed DNA methylation, which is found in the oldest nematode clades (Clades I and II). It is also found in plants and fungi, but has been lost in most animals. This finding begs the intriguing question: Might this be the ancestral mechanism of transposon silencing in animals?
“It’s completely unprecedented and shocking to see so many independent losses of the piRNA pathway across a single phylum. There are no other examples of that in the animal kingdom that we know of,” says Peter Sarkies from the CSC’s Transgenerational Epigenetic Inheritance and Evolution Group. If we can understand the selective forces that led to piRNAs being lost, says Sarkies, we might be able to get some insight into why transposable element proliferation is something that occurs more frequently in cancer. In addition, we might be able to design new treatment against parasitic nematodes, such as filarial nematodesresponsible for elephantiasis and river blindness, he adds.
The work was carried out in collaboration with, among others, Eric Miska from the University of Cambridge, and Murray Selkirk from Imperial College London.
Peter Sarkies, Murray E. Selkirk, John T. Jones, Vivian Blok, Thomas Boothby,
Bob Goldstein, Ben Hanelt, Alex Ardila-Garcia, Naomi M. Fast, Phillip M. Schiffer,
Christopher Kraus, Mark J. Taylor, Georgios Koutsovoulos, Mark L. Blaxter, Eric A. Miska: Ancient and Novel Small RNA Pathways Compensate for the Loss of piRNAs in Multiple Independent Nematode Lineages, PLOS Biology, February 10, 2015, DOI:10.1371/journal.pbio.1002061
Almut Caspary
Institute of Clinical Science
Faculty of Medicine



 Synergy: Leverhulme Trust Research Project Grant awarded to CSC and Imperial College Researchers
Synergy: Leverhulme Trust Research Project Grant awarded to CSC and Imperial College Researchers